Imaging single mRNA molecules in live cells has proved to be very useful in studying mRNA localization, as well as mRNA transcription. Specifically, by using the MS2-like systems, it is possible to follow the synthesis of single mRNAs, thus determining the rate of transcription.

Tagging an mRNA with MS2-like systems allows following the transcription of that gene by imaging. Adapted from Larson et al. (2011)
By putting the MBS (MS2-coat protein binding sequence) or PBS ((PP7-coat protein binding sequence) at the 5’UTR of a gene, one can follow the fluctuation of fluorescence at the transcription site. In essence, the MS2-tagged mRNA should appear as a bright spot in the nucleus as soon as the MBS sequence is transcribed. Once the mRNA is fully processed and exported from the nucleus, the fluorescence signal of the transcription site should decrease.
What happens if you have the same gene being transcribed simultaneously by multiple polymerases?
Well if the MBS is at the 3’UTR, then the duration of the MS2 signal at the transcription site for each mRNA should be short (the time it takes to complete synthesis of the 3’UTR and the 3’-end processing).
However, if the MBS is at the 5’UTR, then the signal lasts from early elongation until 3’-end processing and export. For the GLT1 gene in yeast, it is ~200 seconds. Since that gene is ~6400 bases long, than we can calculate that the rate of RNA polymerase is less than 32 bases per second (we do not know the 3’-end processing/termination time).

Time-dependent activity of individual reporter genes (PP7-GFP, green; Nup49-tomato, red). At time t = 0 min, both cells show a TS near the periphery of the nucleus. At t = 2 min, an additional TS corresponding to the duplicated gene has turned on in the upper cell (white and blue arrows). Right: Intensity trace (green line) of the transciption site. The gray line is the intensity of a cytosolic mRNA; the black line is a background intensity at an arbitrary position in the nucleus. Adapted from Larson et al (2011).
We can calculate the real elongation rate by subtracting the signal dwelling time (how long we see the fluorescent spot) with the 3’UTR MBS from the dwelling time with the 5’UTR. Using this method it was possible to calculate the elongation rate of RNA polymerase II on the MDN1 gene in yeast (it is the longest gene in yeast, almost 15,000 bases long). The rate was found to be 20±8 bases per second. Knowing that, we can subtract the time needed for the remaining of the 3’UTR downstream of the 3’UTR MBS, and what is left is the termination time. For the MDN1 gene it is 70 seconds (i.e. it takes the 3’-end processing and export machinery more than a minute to get the mRNA away from the nucleus).
However, this method requires measuring different genes (one with 5’UTR-MBS and another with 3’UTR-MBS) in different cells, getting their average, then subtracting. It would be much more accurate to measure a single gene in a single cell and get an absolute value from that measurement.
Here comes the development of two-color system. Instead of using just MBS, we insert the MBS at the 3’UTR and the PBS at the 5’UTR. We then express PCP-GFP and MCP-RFP and we can have a two colored mRNA. Upon expression, the GFP signal comes first and the RFP signal comes later. By measuring the duration from appearance of GFP to appearance of RFP, we can directly get the elongation rate of a single RNA polymerase enzyme, on a single gene, in a single cell.
That is exactly what Sami from our lab did. He tagged the MDN1 gene with the PBS and MBS and measured the elongation rate in single cells. On average, the rate was found to be 25 bases per second. This is agreeable with the results I mentioned above.
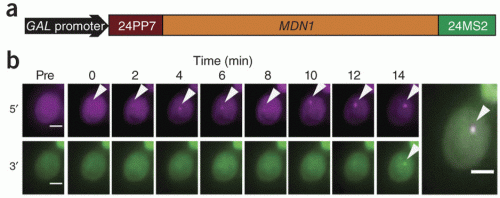
(a) Schematic showing the tagging of MDN1 gene wirh 5′ PP7 loops and 3′ MS2 loops. (b) Appearance of 5′ (magenta) and 3′ (green) transcription site signals at indicated times after induction. Final frames show an overlay of both signals. Adapted from Hocine et al. (2012)
But more interesting was the high variability between cells. The elongation rates of the same gene in differnt cells ranged from as slow as 14 bases per second to the fast rate of 61 bases per second.
What does it mean? That the text-book view that transcription elongation is constant is not true. In recent years it was shown by different methods that transcription elongation is regulated, and that RNA polymerase II is prone to pausing. Here is direct evidence that indeed the same gene is elongated at different rates in different cells. Which means that we still have a lot to learn about transcription in general, and elongation in particular.
![]() Larson DR, Zenklusen D, Wu B, Chao JA, & Singer RH (2011). Real-time observation of transcription initiation and elongation on an endogenous yeast gene. Science (New York, N.Y.), 332 (6028), 475-8 PMID: 21512033
Larson DR, Zenklusen D, Wu B, Chao JA, & Singer RH (2011). Real-time observation of transcription initiation and elongation on an endogenous yeast gene. Science (New York, N.Y.), 332 (6028), 475-8 PMID: 21512033
Hocine S, Raymond P, Zenklusen D, Chao JA, & Singer RH (2012). Single-molecule analysis of gene expression using two-color RNA labeling in live yeast. Nature methods PMID: 23263691








Hi! very suddenly, I found your blog and I must say, it’s great! I have a question about the MCP you normally use in your lab; is it the wild type version or mutant? Does is really make a differenc in terms of results?
LikeLike
Pingback: Imaging of miRNA-mediated translational repression and mRNA decay at single molecule resolution | greenfluorescentblog